Plk1のPBDに依存した結合阻害剤の探索
概要
細胞内のタンパク質は翻訳後修飾を受けることで、他のタンパク質との相互作用状態が変化するものが少なくない。翻訳後修飾の中で、リン酸化はその代表的なものであり、特定の残基がリン酸化されることによって他のタンパク質と結合できるようになるタンパク質が多く知られている。この結合を阻害する物質はリン酸化依存結合の生理的意義を解析するために有用であるのみならず、リン酸化依存結合を介した細胞増殖・分化を調節する分子標的薬としての利用も可能である。標的タンパク質にリン酸化依存結合するPlk1のポロボックスドメイン(PBD)依存結合を測定し、結合を阻害する小分子をハイスループットに探索することが可能である。
|
|
|
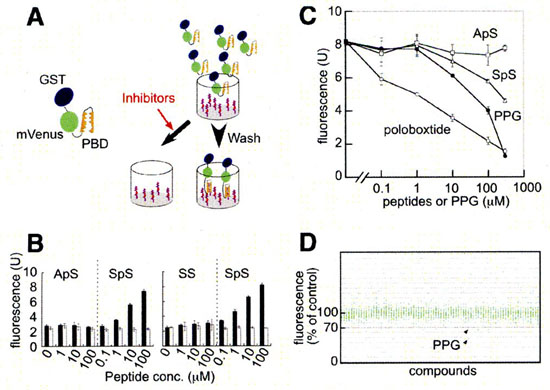
Screening system for inhibitors of PBD-dependent binding and identification of PPG as an inhibitor of PBD-dependent binding.
A. Schematic representation of the screening system. Target phosphopeptides for PBD were covalently bound in 96-well plates, and binding of GST-mVenus-PBD to the phosphorpeptides was quantitated by spectrofluorometry. When inhibitors are present, PBD will fluoresce less. B. Phosphopeptides of PBD-binding sequence (SpS; C-EEEGFGSSpSPVKSPAAP) and their mutated (ApS) or unphosphorylated versions (SS) were bound to 96-well plates using peptides at concentrations as indicated. Binding of wild-type (solid bars) or H538A/K540M mutant forms (open bars) of PBD was determined by spectrofluorometry. C. Effects of various concentrations of phosphopeptides encoding optimal PBD-binding sequence (poloboxtide; MAGPMQSpTPLNGAKK), SpS (C-EEEGFGSSpSPVKSPAAP) and ApS (C-EEEGFGAApSPVKSPAAP) on PBD-dependent binding. Effects of same concentrations of PPG were also examined. In this particular experiment, about 2,000 fluorescence unit of GST-mVenus-PBD were mixed with each competitors, put into each well, washed and bound fluorescence units were plotted. Each symbol represents mean value, while each vertical line indicates standard deviation (n=3). D. Results of initial screening of about 2,500 compounds. The effect of each compound (at 50 μM) on the PBD-dependent binding was standardized and represented as percentage value of control (without compound; ie. 100% means no inhibition). The effects of more than 90% compounds were within the range of 100 ± 20%, only PPG inhibited the binding more than 30%, reproducibly (arrows). (Ref:1)
|
References:
-
N. Watanabe, T. Sekine, M. Takagi, J.I. Iwasaki, N. Imamoto, H. Kawasaki,H. Osada.
Deficiency in chromosome congression by the inhibition of PLK1 polo box domain-dependent recognition.
J. Bio. Chem., 284, 2344-2353 (2009).